The first caffeine-'addicted' bacteria
Some people may joke about living on caffeine, but scientists now have genetically engineered E. coli bacteria to do that—literally. Their report in the journal ACS Synthetic Biology describes bacteria being "addicted" to caffeine in a way that promises practical uses ranging from decontamination of wastewater to bioproduction of medications for asthma.
Jeffrey E. Barrick and colleagues note that caffeine and related chemical compounds have become important water pollutants due to widespread use in coffee, soda pop, tea, energy drinks, chocolate and certain medications. These include prescription drugs for asthma and other lung diseases. The scientists knew that a natural soil bacterium, Pseudomonas putida CBB5, can actually live solely on caffeine and could be used to clean up such environmental contamination. So they set out to transfer genetic gear for metabolizing, or breaking down, caffeine from P. putida into that old workhorse of biotechnology, E. coli, which is easy to handle and grow.
The study reports their success in doing so, as well as use of the E. coli for decaffeination and measuring the caffeine content of beverages. It describes development of a synthetic packet of genes for breaking down caffeine and related compounds that can be moved easily to other microbes. When engineered into certain E. coli, the result was bacteria literally addicted to caffeine. The genetic packet could have applications beyond environmental remediation, the scientists say, citing potential use as a sensor to measure caffeine levels in beverages, in recovery of nutrient-rich byproducts of coffee processing and for the cost-effective bioproduction of medicines.
More information: Decaffeination and Measurement of Caffeine Content by Addicted Escherichia coli with a Refactored N-Demethylation Operon from Pseudomonas putida CBB5, ACS Synth. Biol., DOI: 10.1021/sb4000146
Abstract
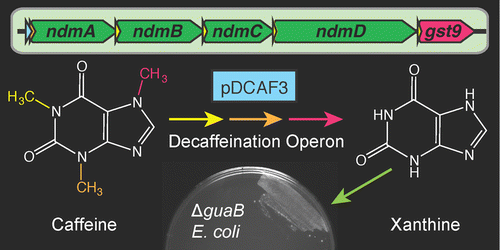
The widespread use of caffeine (1,3,7-trimethylxanthine) and other methylxanthines in beverages and pharmaceuticals has led to significant environmental pollution. We have developed a portable caffeine degradation operon by refactoring the alkylxanthine degradation (Alx) gene cluster from Pseudomonas putida CBB5 to function in Escherichia coli. In the process, we discovered that adding a glutathione S-transferase from Janthinobacterium sp. Marseille was necessary to achieve N7-demethylation activity. E. coli cells with the synthetic operon degrade caffeine to the guanine precursor, xanthine. Cells deficient in de novo guanine biosynthesis that contain the refactored operon are ″addicted″ to caffeine: their growth density is limited by the availability of caffeine or other xanthines. We show that the addicted strain can be used as a biosensor to measure the caffeine content of common beverages. The synthetic N-demethylation operon could be useful for reclaiming nutrient-rich byproducts of coffee bean processing and for the cost-effective bioproduction of methylxanthine drugs.
Journal information: ACS Synthetic Biology
Provided by American Chemical Society