Researchers develop new interferometric single-molecule localization microscopy
Various image-based central position estimation (termed "centroid fitting") methods such as 2-D Gaussian fitting methods have been commonly used in single-molecule localization microscopy (SMLM) to precisely determine the location of each fluorophore. Yet it is still a challenge to improve the single-molecule lateral localization precision to molecular scale (< 2 nm) for high throughput nanostructure imaging.
In a study published online in Nature Methods, Prof. Xu Tao and Prof. Ji Wei from the Institute of Biophysics of the Chinese Academy of Sciences developed a new interferometric single-molecule localization microscopy process with fast modulated structured illumination, called Repetitive Optical Selective Exposure (ROSE).
ROSE utilizes six different direction and phase interference fringes to excite the fluorescent molecules. The intensity of the fluorescent molecules is closely related to the phase of the interference fringes. A fluorescence molecule is located by the intensities of multiple excitation patterns of an interference fringe, providing around two-fold improvement in localization precision. This technique has pushed the resolution of single-molecule localization microscopy (SMLM) to less than 3 nm (~1 nm localization precision).
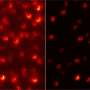
The researchers first designed three different lattice grids of DNA origami structures with 5-, 10- and 20-nm point-to-point spacing to verify the performance of ROSE. Both conventional centroid fitting and ROSE could resolve the 20-nm structure at the same photon budget. ROSE could also clearly resolve the 10-nm distance, which could not be resolved by centroid fitting.
The researchers demonstrated that ROSE could resolve a 5-nm structure at a resolution of ~3 nm over a large field of view of 25 x 25 μm2, which means that ROSE has the ability to push the resolving power of SMLM to the molecular scale.
In addition, using ROSE to analyze cellular nanostructures, the researchers showed that ROSE has advantages in resolving the hollow structure of single microtubule filaments, small clathrin-coated pits (CCPs) and cellular nanostructures of actin filament. The Fourier ring correlation (FRC) analysis indicated that ROSE improved final resolution by twofold compared with the centroid fitting method.
ROSE can be extended to 3-D nanometer-scale imaging by introducing additional excitation patterns along the axial direction. The researchers envision that this method could extend the application of SMLM in biomacromolecule dynamic analysis and structural studies at the molecular scale.
More information: Molecular resolution imaging by repetitive optical selective exposure, Nature Methods (2019). DOI: 10.1038/s41592-019-0544-2 , nature.com/articles/s41592-019-0544-2
Journal information: Nature Methods
Provided by Chinese Academy of Sciences