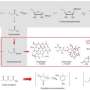
Helping robots to build new antibiotics
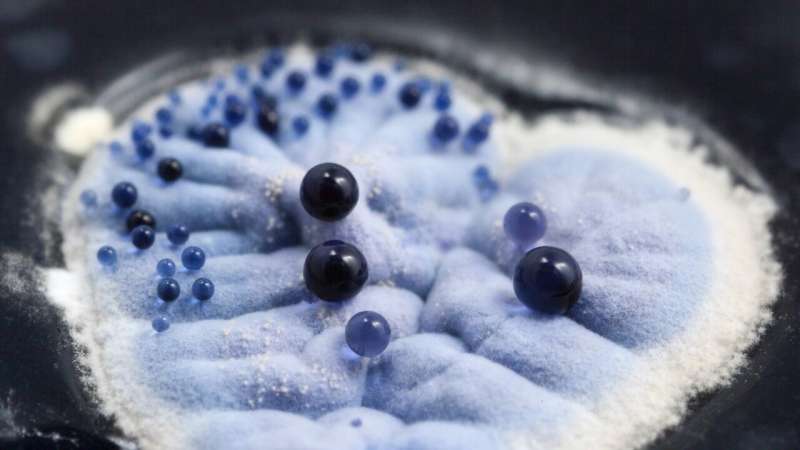

A team from The University of Manchester have engineered a common gut bacterium to produce a new class of antibiotics by using robotics. These antibiotics, known as class II polyketides, are also naturally produced by soil bacteria and have antimicrobial properties which are vital in the modern pharmaceutical industry to combat infectious diseases and cancer.
The naturally produced Escherichia coli bacteria are difficult to work with as they grow in dense clumps that are incompatible with the automated robotic systems used for modern biotechnology research. By transferring the production machinery from the soil bacteria into E. coli, the Manchester team is now making this class of antibiotics accessible for much more rapid exploration.
This breakthrough could be vital for the ongoing combat against antimicrobial resistance, as recently developed automated robotics systems can now be used to create new antibiotics in a fast and efficient way.
In this work, published in the journal PLOS Biology, the group led by Professor Takano at The University of Manchester show the potential of this approach. By combining the bacterial production machinery with enzymes from plants and fungi, it was possible to produce new chemical compounds not previously seen in nature. Using this plug-and-play platform, it will now be possible to explore and engineer polyketides using robotic systems to develop new and diversified polyketides in an automated, rapid and versatile fashion.
Eriko Takano, Professor of Synthetic Biology said: "Nature is a huge treasure trove for powerful chemical compounds to treat a wide range of diseases. However, the most interesting chemicals often come from organisms that are difficult to work with in the laboratory.
"This has been a major bottleneck for our work on type II polyketides, a group of important chemicals, which are mostly produced by soil bacteria and other microorganisms that are challenging to grow. By successfully moving the production machinery for these compounds into the "laboratory workhorse" bacterium E. coli, we can finally produce and engineer type II polyketides in our rapid robotic systems.
"This not only allows us to trial new polyketides in an automated manner, but we will also be able to quickly rewrite the DNA sequences of the antibiotic biosynthesis pathways and combine them with new components from other organisms, such as medicinal plants and fungi, to produce variations on the antibiotic theme—including compounds that are not produced by the natural pathways, but may have enhanced or novel activities in the treatment of important diseases."
It could take a person a whole year to make and test ten new potential antibiotics, but this automated robotic system can make thousands in that time. This would hugely decrease the time it takes for new antibiotics to reach patients, and provide the necessary agility to react to new pathogen strains and outbreaks.
More information: Matthew Cummings et al. Assembling a plug-and-play production line for combinatorial biosynthesis of aromatic polyketides in Escherichia coli, PLOS Biology (2019). DOI: 10.1371/journal.pbio.3000347
Journal information: PLoS Biology
Provided by University of Manchester