Gene transcription machinery constrains DNA movements, study suggests
Researchers in Japan have discovered that the DNA inside human cells moves around less when its genes are active. The study, which will be published March 1 in the Journal of Cell Biology, suggests that RNA polymerase II (RNAPII)—the key enzyme required to produce messenger RNA molecules from active genes—restricts the movement of DNA by organizing it into a network of interconnected domains.
To fit inside the nucleus of the cell, DNA is organized into chromatin, in which the strands of DNA are wrapped around groups of histone proteins, like thread around a spool, to form structures known as nucleosomes. Nucleosomes can then be folded up into even more compact structures. When a gene is activated, however, its chromatin is thought to open up and, at the same time, become more mobile and dynamic, so that RNAPII can transcribe the gene into messenger RNAs.
Kazuhiro Maeshima and colleagues at the National Institute of Genetics in Mishima, Japan, were therefore surprised when they discovered that the chromatin in human cells becomes more mobile when RNAPII and gene transcription are inhibited.
Maeshima's group used a high-resolution microscopy technique that allowed them to track the movements of individual nucleosomes inside living cells. When the researchers depleted RNAPII from cells, or added drugs that inhibit the enzyme, nucleosomes in the genome clearly became more dynamic, suggesting that RNAPII usually restricts global chromatin movements.
RNAPII and gene transcription activity are naturally reduced when cells enter a dormant state called quiescence or when their DNA is damaged by ultraviolet light. Accordingly, Maeshima and colleagues saw that chromatin was more dynamic in quiescent or UV-irradiated cells. The researchers speculate that these increased movements may help chromatin recruit factors required to repair DNA or restart gene transcription when quiescent cells are reactivated.
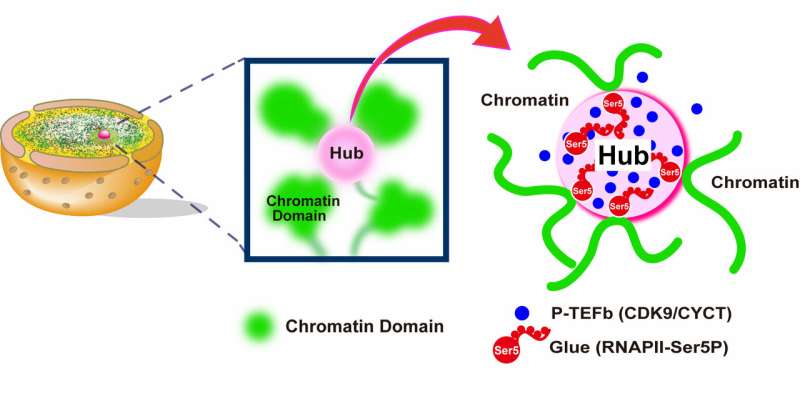
But how does gene transcription affect the global mobility of chromatin when, at any given moment, RNAPII is only transcribing a small fraction of the genome? Based on computer simulations, Maeshima and colleagues propose that clusters of RNAPII and associated factors can bind and connect distant chromatin regions, linking them together in an organized network. These connections are lost when RNAPII is inactivated, breaking up the network and allowing chromatin to become more mobile.
"Our imaging and computational modeling results suggest that chromatin is globally stabilized by loose connections through transcriptionally active RNAPII," Maeshima says. "Our model is compatible with the classical idea of stable transcription factories containing RNAPII, as well as with recent reports that RNAPII and other factors undergo a phase separation process to form dynamic clusters within the nucleus."
More information: Nagashima et al., 2019. J. Cell Biol. DOI: 10.1083/jcb.201811090
Journal information: Journal of Cell Biology
Provided by Rockefeller University Press


















