Study demonstrates 'tunability' of a molecular chaperone

For decades, molecular biologists studying a class of molecular chaperones known as heat shock proteins (Hsp70s) have relied on the Hsp70s found in bacteria as the model system. Now one of the world's experts on the molecule and her team report that their investigation into whether Hsps from mammalian cells behave like those in bacteria reveals "key evolutionary variations" between them.
Lila Gierasch, an expert on Hsp70s at the University of Massachusetts Amherst, with her research team, report that Hsp70s from mammalian cells behave quite differently from bacterial Hsp70s. Because of the important roles Hsp70s play in protein misfolding diseases such as cancer and neurodegenerative diseases, the new findings "will have a major impact on how we think about Hsp70s," she says.
As Gierasch points out, "We've relied on the bacterial version of Hsp70s to study for so long, we thought it was time to ask if eukaryotic Hsp70s behave like those in bacteria or not. After all, it's not too surprising that they might be different because bacteria are so streamlined and have less functional complexity than eukaryotes." Molecular chaperones help cells to maintain healthy proteins by assisting newly synthesized proteins to fold to their functional structures and by protecting cells from stresses like heat shock, which damage proteins, she adds.
"I want to emphasize that what we learned in bacteria is absolutely essential to understanding the more sophisticated mammalian chaperone family members. We have dissected the architecture of the bacterial Hsp70 and related it to its functional structural changes. We knew the importance of key interfaces between functional domains. We noted that there were widespread evolutionary variations in these interfaces in the mammalian Hsp70s. We postulated that these variations would be reflected in functional diversification."
Details of this work funded by the NIH's Maximizing Investigators' Research Awards program appear this week in Proceedings of the National Academy of Sciences. Gierasch's co-authors include postdoctoral researcher Wenli Meng, research assistant professor Eugenia Clerico and an undergraduate, Natalie McArthur, now a graduate student at Columbia.
Gierasch explains that the versatile chaperone molecules, known as universal tools of cellular protein folding, interact with many different types of protein and are involved in many cellular functions. Hsp70s help proteins to fold, to translocate across membranes, to assemble into complexes, to be targeted for degradation, and to avoid harmful misfolding and aggregation. They are thought of as hubs in the cell's finely balanced protein quality control network for good reason, she notes.
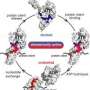
The researchers point out that Hsp70s accomplish these many and varied functions by a conserved mechanism that relies on cycles of nucleotide-modulated binding and release of their client proteins, a process Gierasch refers to as "domain docking and undocking." To examine the cycles of domain docking and undocking of both eukaryotic and bacterial Hsps in fine detail, Gierasch and colleagues used domain dissection techniques, biochemical assays and specialized nuclear magnetic resonance spectrometry experiments.
They report finding "significant differences" between the workings of the bacterial and eukaryotic chaperones, notably that the bacterial Hsp70 favors a state in which the two domains are "intimately docked significantly more" compared to the more loosely bound eukaryotic chaperones. Gierasch says, "In the bacterial cell, the chaperone may hold onto its client longer. Imagine hands holding a rope. In the eukaryotic cell it looks like the hand is grabbing transiently and letting go all the time, while in the bacterial cell the molecule is holding on tight most of the time."
The molecular biologist speculates that it might be evolutionarily advantageous for eukaryotic cells to have developed a more flexible binding technique that is open to handing its clients off for downstream processes more quickly and smoothly. "It may be that the bacterial function is more specific and narrower, dominated by biosynthesis of protein and providing assistance in folding. But the eukaryotic Hsp70 may be required to pass its client on to partners in any of the array of functions it is participating in—the Hsp70 should not hold too tight. If the client dwells in one Hsp70 for too long, it won't be handed off to the next process," Gierasch points out.
"These results underline the tunability of Hsp70 functions by modulation of allosteric interfaces through evolutionary diversification," the authors state, "and also suggest sites where the binding of small-molecule modulators could influence Hsp70 function." These insights should help researchers understand the mechanism of Hsp70 functional diversities and design specific small-molecule Hsp70 modulators, they add.
Being able to "tune" Hsp70s has long been a goal of medical researchers seeking ways to treat diseases such as cancer and neurological disorders. As Gierasch explains, however, the chaperone molecules are so intimately involved with so many cell processes that attempting to modulate any one of them is going to affect other processes.
"If you want to cure cancer you might want to inhibit Hsp70s," she notes, "but if you want a therapy for Alzheimer's, which is a protein-folding disease, you want to activate them. Our new deeper understanding of the eukaryotic Hsp70s may offer a route to modulating them with more specificity. It may give us the ability to isolate and regulate a particular function."
More information: Wenli Meng et al, Allosteric landscapes of eukaryotic cytoplasmic Hsp70s are shaped by evolutionary tuning of key interfaces, Proceedings of the National Academy of Sciences (2018). DOI: 10.1073/pnas.1811105115
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Massachusetts Amherst