Self-assembling protein filaments designed and built from scratch
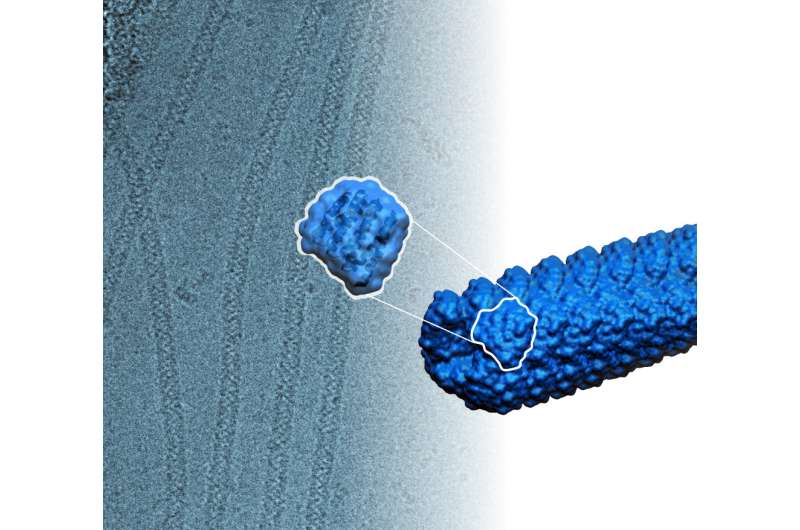
For the first time, scientists have created, from scratch, self-assembling protein filaments.
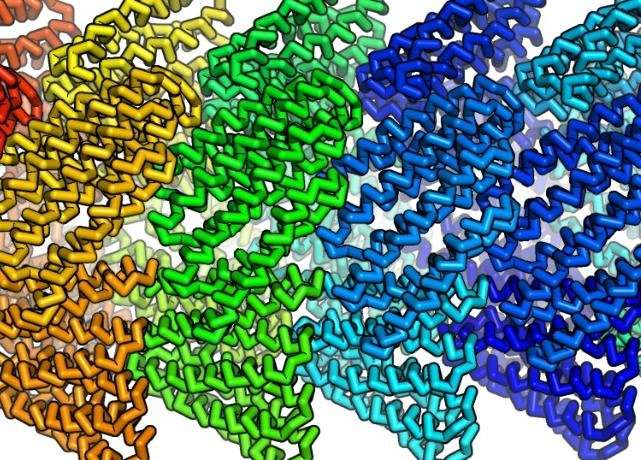
These were constructed from identical protein subunits that snap together spontaneously to form long, helical, thread-like structures.
In the natural world, protein filaments are essential components of several structural and moving parts in living cells, as well as many body tissues.
These include the cytoskeletons that give cells their shape, the cellular microtubules that orchestrate cell division, and the most common protein in our bodies, collagen, which gives both strength and flexibility to our cartilage, skin and other tissues.
"Being able to create protein filaments from scratch—or de novo—will help us better understand the structure and mechanics of naturally occurring protein filaments and will also allow us to create entirely novel materials unlike any found in nature," said David Baker, a professor of biochemistry at the University of Washington School of Medicine, and director of the UW Institute for Protein Design, who directed the project. He is also a Howard Hughes Medical Institute investigator.
Such materials might include man-made fibers that equal or surpass the strength of spider silk, which by weight is stronger than steel, Baker said. He also mentioned the possibility of nano-scale wire circuitry.
To design the filaments, the researchers used a computer program developed in the Baker laboratory, called Rosetta, that can predict the shape of a protein from its amino acid sequence.
To function properly, proteins must fold into a precise shape. This folding is driven by properties of the individual amino acids and how they interact with each other and the surrounding fluid environment. The forces of attraction and repulsion drive the protein to come to rest in a shape that has the lowest energy level.
By calculating which shape would balance out these forces of attraction and repulsion to yield the lowest total energy level, Rosetta can predict, with a high degree of accuracy, the shape a protein will assume in nature.
Using Rosetta, the researchers set out to design small proteins that had amino acids on their surface that would cause them to latch onto each other. This allowed them to assemble into a helix by aligning like steps in a winding staircase. For the helix to be stable, the designed protein binds other copies positioned above and below it as the helix winds around, tier on tier.
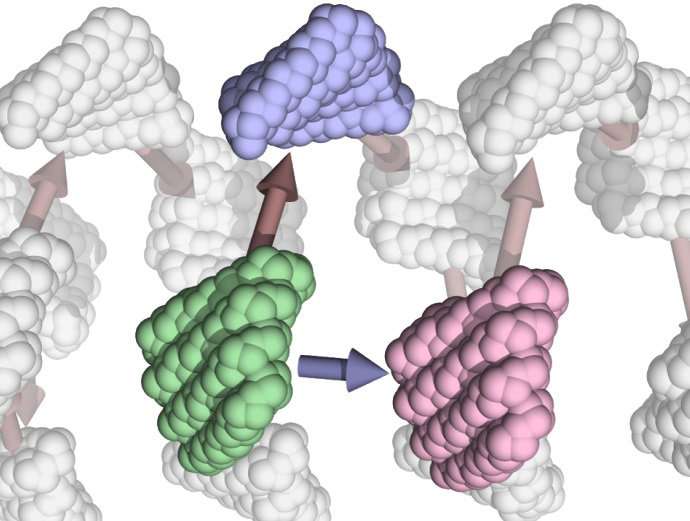
"We were eventually able to design proteins that would snap together like Legos," said Hao Shen, a Ph.D. candidate at the UW Molecular Engineering & Sciences Institute. He and Jorge Fallas, an acting instructor in biochemistry at the UW School of Medicine, are lead authors of a paper describing the approach.
This paper will be published online by the journal Science on Thursday, Nov. 8, 2018.
Fallas said that the designed proteins are relatively small. They are made up of about only 180 to 200 amino acids and measure only about one nanometer in length, but assemble into stable filaments more than 10,000 nanometers long. A nanometer is 1 billionth of a meter, or about the width of 10 hydrogen atoms lined up side by side.
The researchers also showed that that, by tinkering with the designed protein's concentration in solution and by adding caps that inhibited the design's ability to bind, they could drive the filaments to grow or to disassemble.
"The ability to program the dynamics of filament formation will give us insights into how filament assembly and disassembly is regulated in nature," said Baker. "The stability of these proteins suggest they could serve as easily modifiable scaffolds for a range of applications ranging from new diagnostic tests to nano-electronics."
More information: "De novo design of self-assembling helical protein filaments" Science (2018). science.sciencemag.org/cgi/doi … 1126/science.aau3775
Journal information: Science
Provided by University of Washington