Bioengineers create pathway to personalized medicine
Engineering cellular biology, minus the actual cell, is a growing area of interest in biotechnology and synthetic biology. It's known as cell-free protein synthesis, or CFPS, and it has potential to provide sustainable ways to make chemicals, medicines and biomaterials.
Unfortunately, a long-standing gap in cell-free systems is the ability to manufacture glycosylated proteins—proteins with a carbohydrate attachment. Glycosylation is crucial for a wide range of important biological processes, and the ability to understand and control this mechanism is vital for disease treatment and prevention.
Matthew DeLisa, the William L. Lewis Professor of Engineering in the Smith School of Chemical and Biomolecular Engineering at Cornell University, and Michael Jewett, associate professor of chemical and biological engineering at Northwestern University, have teamed up on a novel approach that bridges this gap. Their system, the first of its kind, capitalizes on the recent advances in CFPS while adding the crucial glycosylation component in a simplified, "one-pot" reaction. The protein of choice could then be freeze-dried and reactivated for point-of-use synthesis by simply adding water.
DeLisa and Jewett are co-senior authors of "Single-pot Glycoprotein Biosynthesis Using a Cell-Free Transcription-Translation System Enriched with Glycosylation Machinery," published July 12 in Nature Communications.
Thapakorn Jaroentomeechai, Ph.D. student in the DeLisa Research Group, and Jessica Stark '12, Ph.D. student in the Jewett group, are co-first authors.
"If you really want to have a useful, portable and deployable vaccine or therapeutic protein technology that's cell free, you have to figure out the carbohydrate attachment," DeLisa said. "That's, in essence, what we've done in a very powerful way."
This work could impact development of decentralized manufacturing strategies. Rapid access to protein-based medicines in remote settings could change lives; new biomanufacturing paradigms suitable for use in low-resource settings might promote better access to costly drugs through local, small-batch production.
DeLisa has done a great deal of research on the molecular mechanisms underlying protein biogenesis in the complex environment of a living cell, such as Escherichia coli (E. coli). While his lab has made some notable breakthroughs, the limitations of this area, he said, are the cell walls themselves.
Jewett's lab at Northwestern has invested much of its research efforts into cell-free synthetic biology, which leverages nature's most elegant biomachinery outside the confines of the cell, so a collaboration was a natural extension of both labs' work.
"In bacterial cell engineering, you're constantly in a tug of war," Jewett said. "You're introducing a mechanism or capability that's of interest to you as a scientist, but what the cell is trying to do for itself is grow and survive."
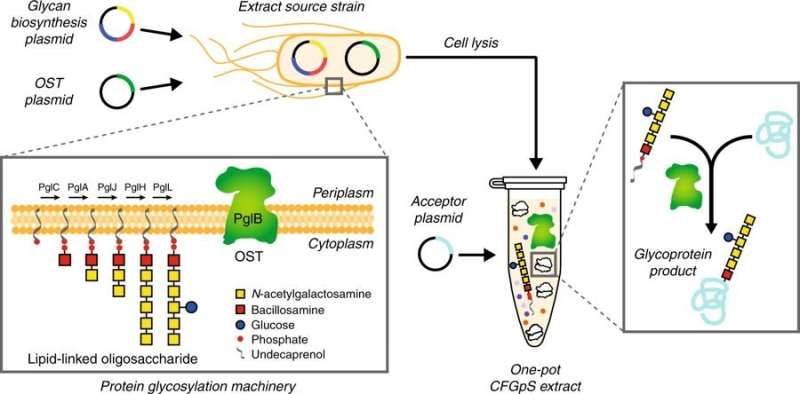
For their new method, the team prepared cell extracts from an optimized laboratory strain of E. coli, CLM24, that were selectively enriched with key glycosylation components. The resulting extracts enabled a simplified reaction scheme, which the team has dubbed cell-free glycoprotein synthesis (CFGpS).
"A major advance of this work is that our cell-free extracts contain all of the molecular machinery for protein synthesis and protein glycosylation," Stark said. "What that means is you only need to add DNA instructions for your protein of interest to make a glycoprotein in CFGpS. This is a drastic simplification from cell-based methods and allows us to make sophisticated glycoprotein molecules in less than a day."
And the CFGpS method is highly modular, allowing for the use of distinct and diverse extracts to be mixed for the production of a variety of glycoproteins.
"Because we chose E. coli, which lacks its own glycosylation machinery, to build our CFGpS platform, it gave us a blank slate for bottom-up engineering of any desired glycosylation system," Jaroentomeechai said. "This gives us the unique ability to control carbohydrate structures and purities of the glycoproteins at levels that are not currently achievable in other cell-based expression systems."
Even in developed countries like the U.S., the move toward personalized medicine makes this type of on-demand drug production protocol attractive. "You could use a test tube instead of a 50,000-liter bioreactor to make your product, which opens the door to a personalized biomanufacturing paradigm where each patient can receive a unique protein medicine tailored to their physiology," he said.
More information: Thapakorn Jaroentomeechai et al, Single-pot glycoprotein biosynthesis using a cell-free transcription-translation system enriched with glycosylation machinery, Nature Communications (2018). DOI: 10.1038/s41467-018-05110-x
Journal information: Nature Communications
Provided by Cornell University