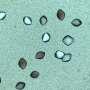
Molecular details of protein crystal nucleation uncovered
A team of researchers led by Dr. Mike Sleutel from the VIB-VUB Center for Structural Biology in collaboration with scientists from the Institute for Complex Molecular Systems of the Eindhoven University of Technology, and the CNRS in Grenoble, have for the first time uncovered the molecular details of protein crystal nucleation, a process with great medical and scientific relevance. The team also developed a new methodology to study a broad class of systems that have remained elusive to date. Their results are published in Nature.
Dr. Mike Sleutel (VIB-VUB): "It will be exciting to see this new technique being applied in the future to follow protein self-assembly processes that are implicated in a range of pathological disorders, such as liquid-liquid phase separation in eye cataract formation or the formation of amyloid fibers associated with a range of neurological disorders."
Protein crystals bear great medical and scientific relevance. For decades, they have been essential for structural biologists to solve the three-dimensional structures of proteins, but protein crystals are also used as bio-pharmaceutical delivery agents. Crystalline suspensions are attractive formulations to store and administer active pharmaceutical compounds because of their long-term shelf life, low solvent viscosity, and slow dissolution rate. Perhaps the best known example is insulin: insulin shots comprise the subcutaneous injection of a suspension of insulin microcrystals which dissolve slowly to yield a steady and sustained delivery over time. Despite their tremendous potential, there are two factors that limit the use of protein crystals in a broad range of applications.
Challenges in developing protein crystals
First, growing protein crystals, as many molecular biologists will say, is more an art than a science. In fact, for many proteins, crystallization can be excruciatingly difficult. This in part follows from the fact that scientists don't understand the early stages of protein crystal formation. Any crystal originates from a nucleus, a tiny crystalline seed, which forms by the spontaneous grouping of a few molecules in solution that have to adopt a regular organization in three-dimensions. How the molecules realize this improbable feat has remained a mystery up until this point.
Secondly, a single protein can crystallize in multiple different crystal forms, this is known as polymorphism. Different crystal polymorphs have different characteristics, with the most notable ones the power to diffract X-rays (crucial for 3D structure determination), and the rate at which it dissolves (crucial for drug delivery). As of yet, it is very difficult to guide the crystallization process to the polymorph of one's liking. Scientists believe that polymorph selection takes place at the stage of nucleation, but no one knows exactly how the mechanism works.
A new way to look at the self-assembly of macromolecules
The group of scientists lead by Dr. Mike Sleutel have used state-of-the-art cryo-transmission electron microscopy (Cryo-TEM) to capture the birth of a protein crystal by visualizing the process of nucleation at molecular resolution.
Dr. Heiner Friedrich explains: "Because the process happens so rapidly, and at such a small length scale we needed to cryogenically arrest the sample at various stages of the process. Once frozen in time, we use a very sensitive electron microscope to visualize the proteins and how they group together to form a nucleus and finally the protein crystal."
By analyzing the Cryo-TEM images taken from a series of samples at constant time intervals, they could start to puzzle together the series of molecular collisions that need to take place to form a crystalline nucleus. Dr. Mike Sleutel continues: "We were struck by the unexpected complexity of the process, which proved to be far more intricate than the working models we and other in the field had prior to these observations. For the protein that we used in our study we uncovered a hierarchical self-assembly process that involves three subsequent stages of self-assembly at ever increasing length scales." These observations are the first of their kind and provide an new way to look at the self-assembly processes of macromolecules into larger structures.
But the team went even one step further, and compared the nucleation pathways of multiple polymorphs. They showed that polymorph selection is dictated by the architecture of the smallest possible fragments formed at early time-points. Once such structures are formed, the faith of the system is set. Dr. Alexander Van Driessche explains: "By analyzing and understanding the differences in structure of the various nuclei, we developed strategies to guide the polymorph selection process. We achieved this by gently tweaking the different modes of interaction that exist between the molecules, steering the nucleation process in the direction of our choosing." The team believes that the new insights and methodology will significantly advance the development of protein crystals for 3D structure determination and medical applications.
More information: Alexander E.S. Van Driessche et al., Molecular nucleation mechanisms and control strategies for crystal polymorph selection, Nature 2018.
Journal information: Nature
Provided by VIB (the Flanders Institute for Biotechnology)