Gender roles in ancient times
Two new studies by Osaka University researchers provide insights on why male and female bodies of the same species differ. The studies show factors that regulate the expression of doublesex1, a gene responsible for the growth of male traits in the ancient crustacean Daphnia magna and the subsequent spatial expression of doublesex1 in embryo development. The studies provide information on how the environment causes genetic changes for gender preference and the evolution of sexual dimorphism.
The rich mane of a lion, the colorful tail of a peacock, examples of sexual dimorphism are abundant. Osaka University Professor Hajime Watanabe has been researching the genes that are the basis for why male and female bodies of the same species differ. In two new papers, his lab reports the molecular regulation and spatial expression of the aptly named gene, doublesex1, in Daphnia magna. This ancient crustacean provides a model for explaining the evolution of sexual dimorphism in the animal kingdom.
"Sex determination can be broadly divided into two categories: genetic sex determination and environmental sex determination (ESD)," says Watanabe.
In normal conditions, Daphnia magna reproduces asexually to only form females. However, in times of high stress, such as a shortage of food, it will apply ESD to also asexually produce males, which will contribute to sexual reproduction.
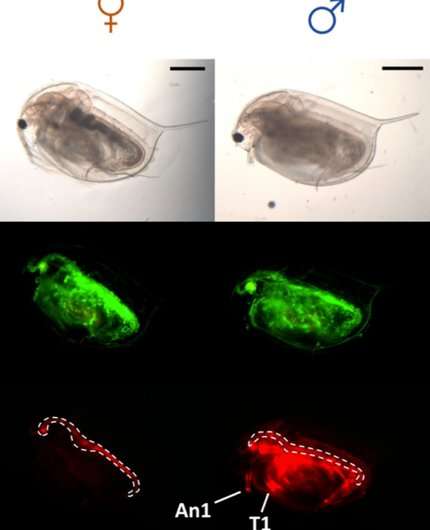
In the first study, using TALEN-based gene editing, the group successfully attached a fluorescent reporter to the doublesex1 gene to watch the spatial expression of doublesex1 in the Daphnia magna embryo in real time. The study shows when and where in the embryo doublesex1 is first expressed and how that expression changes with time to produce male traits.
"Our results suggest a time- and site-specific role of doublesex1. The gene is only recruited when and where it is needed," says Watanabe.
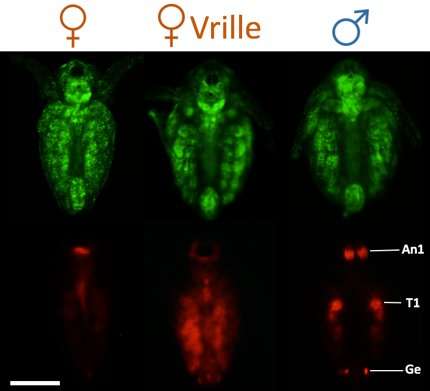
The findings break the expression of dobulesex1 for male development into six stages, including two previously unidentified stages, stomodeal invagination and cumulus migration.
Although both genders carry the gene, the temporal expression of doublesex1 is much longer in males than females, suggesting certain factors behave differently in the two sexes. In the second study, Watanabe demonstrates the transcription factor is responsible for expressing doublesex1. Vrille is known to have a role in growth and circadian rhythms, but the group discovered that it's sensitive to environmental stress. Watanabe's team found that suppressing Vrille expression in male-developing embryos or forcing its expression in female-developing embryos caused the embryos to show signs of the opposite sex and change the doublesex1 expression.
Most of the embryos in the experiments died, probably because Vrille is crucial for many other biological functions besides sex development, but the data, Watanabe says, made it clear that Vrille activates the transcription of doublesex1 through gene co-option.
"Gene co-option is an evolutionary method through which genes take new functions. Humans do not have Vrille. They have an ortholog, E4BP4/NFLIL3," he says, adding that studying this co-option in Daphnia magna could give insight on the evolution of E4BP4/NFLIL3 in humans.
The findings of the studies are consistent with other animals and, Watanabe stresses, support the use of Daphnia magna to study the evolution of sexual dimorphism.
"A number of groups have studied the development of sex-specific traits in model organisms such as mouse and Drosophila. These models are informative, but they are not suitable for studying evolution. Daphnia magna has a more ancestral position and unique sex system," he says.
More information: Quang Dang Nong et al. Mapping the expression of the sex determining factor Doublesex1 in Daphnia magna using a knock-in reporter, Scientific Reports (2017). DOI: 10.1038/s41598-017-13730-4
Nur Syafiqah Mohamad Ishak et al. Co-option of the bZIP transcription factor Vrille as the activator of Doublesex1 in environmental sex determination of the crustacean Daphnia magna, PLOS Genetics (2017). DOI: 10.1371/journal.pgen.1006953
Journal information: Scientific Reports , PLoS Genetics
Provided by Osaka University