Cellular clean-up can also sweep away forms of cancer
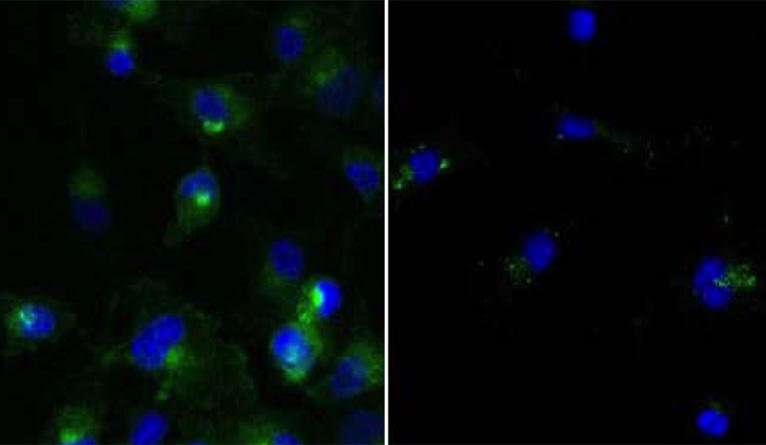
Two new research papers reinforce the benefits of a novel therapy that hijacks the cell's own protein degradation machinery to destroy cancer cells, Yale researchers report Nov. 9 in the journal Cell Chemical Biology.
The new approach to drug discovery, called Proteolysis Targeting Chimeras or PROTACs, was developed in the laboratory of Craig Crews, the Lewis B. Cullman Professor of Molecular, Cellular, and Developmental Biology, professor of chemistry and pharmacology, as well as executive director of the Yale Center for Molecular Discovery.
The system engages the cell's own protein degradation machinery to destroy targeted proteins by tagging them for removal. Most drugs are based on the ability of small molecules to bind to and block the function of disease-causing proteins, but some proteins are resistant to such intervention.
"This system will help us change the current small-molecule drug paradigm that fails to target 75% of rogue proteins," said Crews, scientific founder of Arvinas LLC, the New Haven biotechnology company developing the concept.
The first paper shows for the first time that PROTAC system can target mutant RTK proteins, which have been linked to several forms of cancer. The second paper proves that the PROTAC system can target rogue proteins with greater specificity than traditional approaches.
Yale's George M. Burslem and Blake E. Smith are first authors of the first paper. Smith and Yale's Daniel P. Bondeson are co-first authors of the second paper.
More information: George M. Burslem et al. The Advantages of Targeted Protein Degradation Over Inhibition: An RTK Case Study, Cell Chemical Biology (2017). DOI: 10.1016/j.chembiol.2017.09.009
Daniel P. Bondeson et al. Lessons in PROTAC Design from Selective Degradation with a Promiscuous Warhead, Cell Chemical Biology (2017). DOI: 10.1016/j.chembiol.2017.09.010
Journal information: Cell Chemical Biology
Provided by Yale University