DNA origami: Folded DNA as a building material for molecular devices
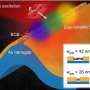
Living things use DNA to store the genetic information that makes each plant, bacterium, and human being unique. The reproduction of this information is made possible because DNA's nucleotides—A's and T's, G's and C's—fit together perfectly, like matching jigsaw puzzle pieces. Engineers can take advantage of the matching between long strands of DNA nucleotides to use DNA as a kind of molecular origami, folding it into everything from nanoscale smiley face artwork to serious drug-delivery devices.
Paul Rothemund discusses the potential of such techniques. Rothemund is research professor of bioengineering, computing and mathematical sciences, and computation and neural systems in the Division of Engineering and Applied Science at Caltech.
What do you do?
I use DNA and RNA as building materials to create shapes and patterns with a resolution of just a few nanometers. The smallest features in the DNA structures we make are about 20,000 times smaller than the pixels in the fanciest computer displays, which are each about 80 microns across. A large part of our work over the last 20 years has been just figuring out how to get DNA or RNA strands to fold themselves into a desired computer-designed shape. As we've mastered the ability to make whatever shape or pattern we desire, we've moved on to using these shapes as "pegboards" for arranging other nano-sized objects, such as protein enzymes, carbon-nanotube transistors, and fluorescent molecules.
Why is this important?
Every task in your body, from digesting food to moving your muscles to sensing light, is powered by tiny nanometer-scale biological machines, all built from the "bottom up" via the self-folding of molecules such as proteins and RNAs. The billions of transistors that make up the chips in our cell phones and computers are tens of nanometers in size, but they are built in a "top down" fashion using fancy printing processes in billion-dollar factories. Our goal is to learn how to build complex artificial devices the way biology builds natural ones—that is, starting from self-folding molecules that assemble together into larger more complex structures. In addition to vastly cheaper devices, this will enable completely new applications, such as man-made molecular machines that can make complex therapeutic decisions and apply drugs only where needed.
How did you get into this line of work?
As an undergraduate at Caltech, I had great difficulty trying to decide how to combine my diverse interests in computer science, chemistry, and biology. Fortunately, the late Jan L. A. van de Snepscheut introduced his computer science class to the hypothetical idea of building a DNA Turing machine—a very simple machine which can nevertheless run every possible computer program. He challenged us, suggesting that someone who knew about both biochemistry and computer science could come up with a concrete way to build such a DNA computer. For a project class in information theory with Yaser Abu-Mostafa, a professor of electrical engineering and computer science, I came up with a pretty inefficient, yet possible, way to do this. At the time, I couldn't interest any Caltech professors in building my DNA computer, but shortly after, USC professor Len Adleman published a paper on a more practical DNA computer in Science. I joined Adleman's lab at USC as a graduate student, and I've been trying to use DNA to build computers or other complex devices ever since. I returned to Caltech as a postdoc in 2001 and became a research professor in 2008.
Journal information: Science
Provided by California Institute of Technology