September 23, 2014 feature
Scientists uncover clues to ATP mystery and how cells work
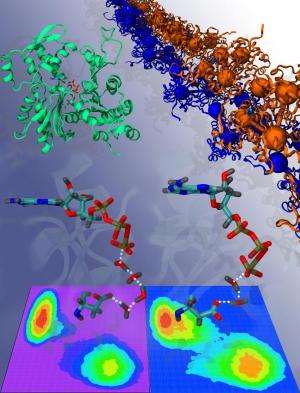
(Phys.org) —Strong, thin fibers called microfilaments, or actin filaments, are present in the cytoplasm of almost every cell in the body. By growing and shrinking, actin filaments play a major role in cellular movement. Since these processes require large amounts of energy, actin filaments use a lot of ATP (adenosine triphosphate), a molecule that is often called the energy currency of the cell.
Although many parts of the cellular movement process are well-understood, there is still some mystery in the details. One mystery in particular is that long polymer chains of actin (called F-actin) have been observed to use ATP at a rate that is more than 40,000 times faster than that of individual monomers of actin (called G-actin). This gigantic rate difference has been experimentally investigated for more than 25 years without a solution.
Now in a new paper in the Journal of the American Chemical Society, researchers Martin McCullagh, Marissa G. Saunders, and Gregory A. Voth at The University of Chicago have used quantum mechanics/molecular mechanics (QM/MM) simulations of G- and F-actin, guided by coarse-grained modeling, to investigate how the structural differences of the two types of actin may contribute to the rate difference at which they use ATP, a process called ATP hydrolysis.
"I think this work provides a fundamental molecular understanding of how actin filaments catalyze ATP hydrolysis, which in turn is critical to their behavior," Voth told Phys.org. "ATP hydrolysis modulates many actin properties, which in turn is very important to the behavior of the cellular cytoskeleton. Some of these properties are relevant to, e.g., the binding of anti-cancer drugs in targeting cancer cells."
In ATP hydrolysis, actin filaments use ATP by breaking off one of its three phosphate atoms. The chemical energy stored in the phosphate bond is released and can be used for many purposes. Here, the energy is used to change the properties of the actin filaments, after which the polymer can be disassembled. This continuous growth and disassembly is what causes cells to move.
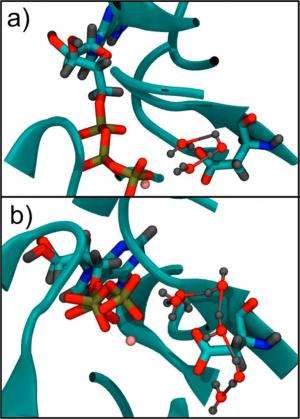
Basically, what the rate difference mystery means is that somehow actin polymerization accelerates ATP hydrolysis. In other words, as the actin forms a filament, the faster it can use ATP. The effect is cyclical, so that the higher the rate of ATP hydrolysis, the higher the rate at which actin can be depolymerized. This activity leads to a remarkable phenomenon called actin "treadmilling," which is essential to cellular movement.
The new QM/MM simulations help explain the dramatic rate increase of ATP hydrolysis that occurs in F-actin compared to G-actin. The simulations show that certain key amino acids change position during polymerization, which promotes the rearrangement of nearby water molecules. The water molecules can rotate more freely, allowing them to easily align into shorter "water wires" that transport protons through the water very quickly—almost instantaneously, in fact. The resulting ease with which the protons can move corresponds to a lowering of the free energy barrier of that process. Since proton transport is a key step in ATP hydrolysis, overall less energy is required for F-actin to hydrolyze the ATP molecules.
The simulations reveal a barrier height reduction that agrees well with the experimentally measured value, supporting the explanation that the favorable proton transport environment in F-actin plays an important role in the increased ATP hydrolysis rate.
"There are many, many proteins that utilize ATP (or GTP) hydrolysis for modulate their behavior," Voth said. "I think this work opens up the possibility to include that process more routinely in the computational study of proteins."
More information: Martin McCullagh, et al. "Unraveling the Mystery of ATP Hydrolysis in Actin Filaments." Journal of the American Chemical Society. DOI: 10.1021/ja507169f
Journal information: Journal of the American Chemical Society
© 2014 Phys.org